Payam Saisan
Dedication
To all the cancer patients, to their courage and love that fuel this work, and to the countless others fighting to defeat cancer.

About Me
My formal background is in Dynamical Systems, Control and Statistical Learning, but my current focus is cancer and neurodegeneration. I work at the intersection of computational biology, machine learning, and control theory toward cures.
I hunt for a rare class of problems — translational inflection points at the intersection of scientific discovery and actionable intervention for patients. This is where fundamental biological insight and data can be analytically harnessed to help engineer therapeutic solutions to otherwise devastating trajectories.
Mathematical Immunology: Bridging AI, Control Theory, & Biology
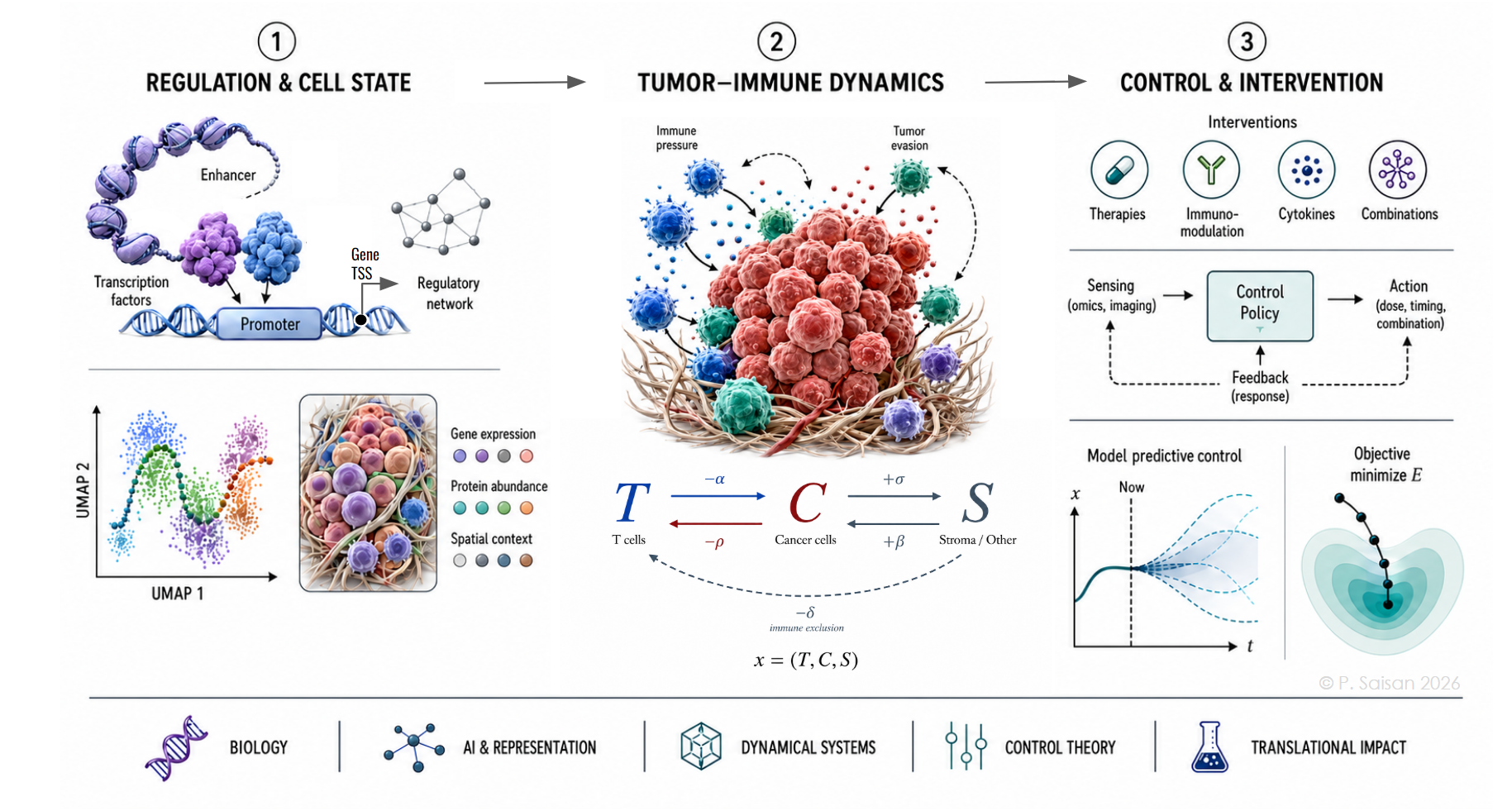
The immune system is one of nature’s most sophisticated adaptive control systems, and possibly the most promising frontier in the battle against cancer. It is a network of programmable molecular agents evolved for distributed adversarial engagement: each immune cell is a molecular dynamical system, collectively giving rise to the astonishing emergent complexity and success of unified immune function.
Evidence increasingly suggests that our best leverage against diseases like cancer may lie within the regulatory programs of our own cells, immune cells in particular: how they can be reprogrammed so that immune dynamics can be modulated toward therapeutic control. I develop predictive and generative models across genomic, cellular, and systems scales to decipher the programming language that governs this system, to help make its reprogramming possible.
Cancer is a cunning asymmetric adversary. It adapts and evades this ultimate defensive barrier from within. Defeating it demands an equally clever asymmetric response at the frontier of biology, AI, control theory, and medicine: to build mechanistic and generative models of the immune system along with cancer and its vulnerabilities, to understand how the immune system can be reprogrammed to change the tide in tumor microenvironments.
I create AI and control-theoretic tools for precision medicine: from regulatory genomics and single-cell/spatial omics to molecular translators in computational pathology and biological state-space control. The mission is to expose cancer’s vulnerabilities and create computational instruments to observe, test, and intervene faster, to save lives.
Let’s Collaborate
If you’re working directly or indirectly on cancer research, feel free to reach out, I reserve time to volunteer and support cancer researchers. Below are a select set of tools reflecting the problems I am working on. Completed tools on GitHub are free to use under their respective licenses. May the force be with you.
📌 Genomics & Gene Regulation
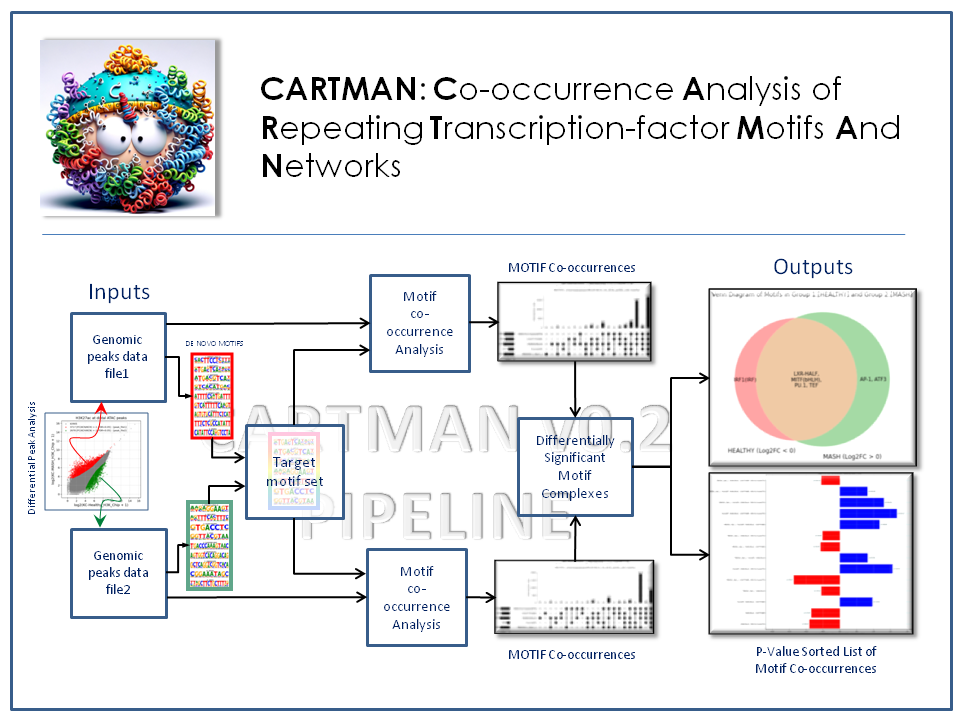 |
🔥 CARTMAN
Co-occurrence Analysis of Repeating Transcription-factor Motifs and Networks. A computational tool for motif discovery and transcription factor co-occurrence analysis in regulatory genomics. 🔗 GitHub Repository |
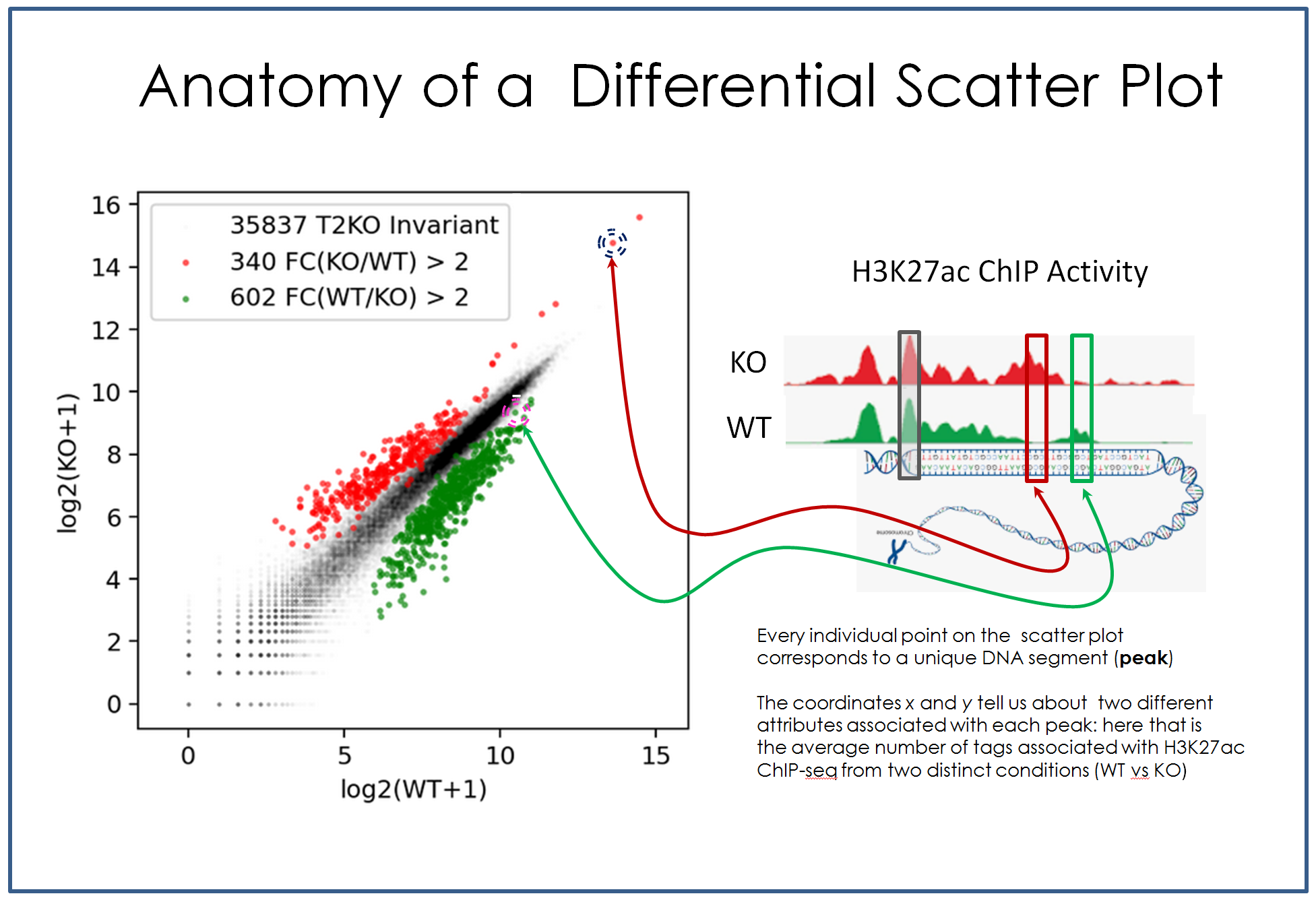 |
🔥 PEAKDIFF
Differential peak analysis for identifying distinct genomic regions between experimental conditions. Includes integration with *ChIP-seq, ATAC-seq*, and *HOMER-based enhancer prediction*. 🔗 GitHub Repository |
 |
🔥 PIPSCOUT
PIPseeker-based Single-Cell Output & UMAP Typing A tool for deciphering PIPseeker's single-cell outputs for downstream analytical pipelines. 🔗 GitHub Repository |
⚙️ Mathematical Modeling & AI for Biological Systems
|
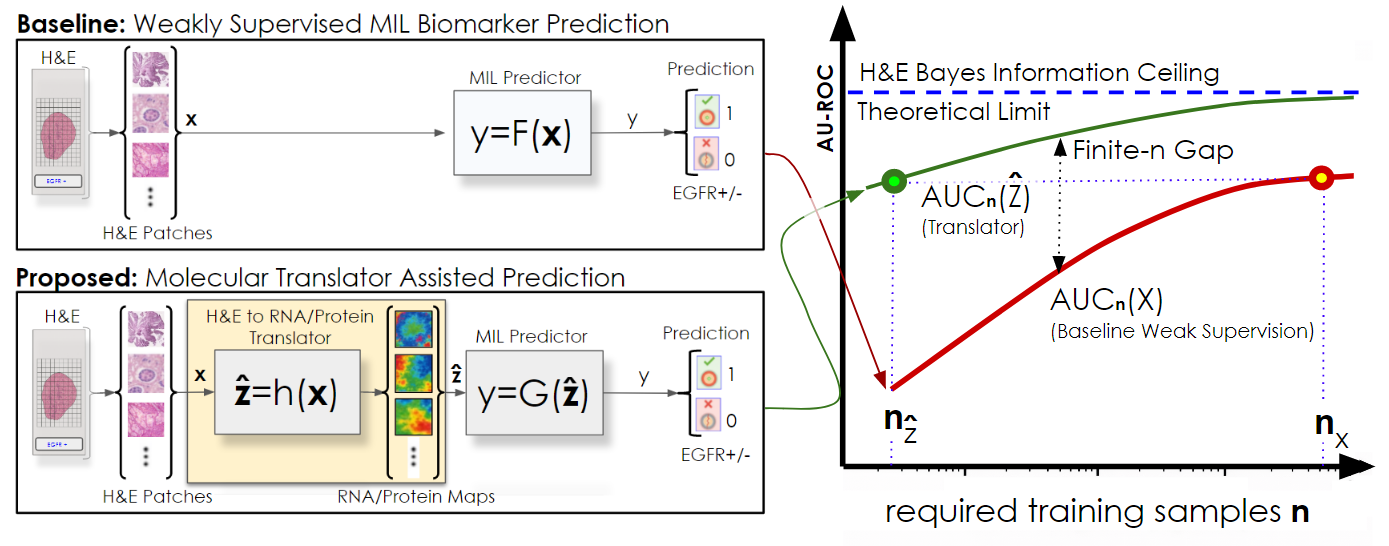
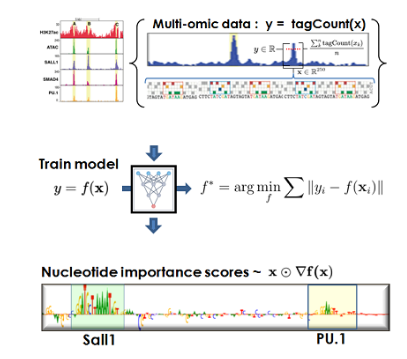
🔥 TRACE in Computational Molecular Pathology: Learning Efficiency of AI Translators under Conserved Information Ceilings. TRACE helps analyze when and how molecular translators can improve biomarker prediction without adding deployment-time information. 🔗 GitHub Repository |
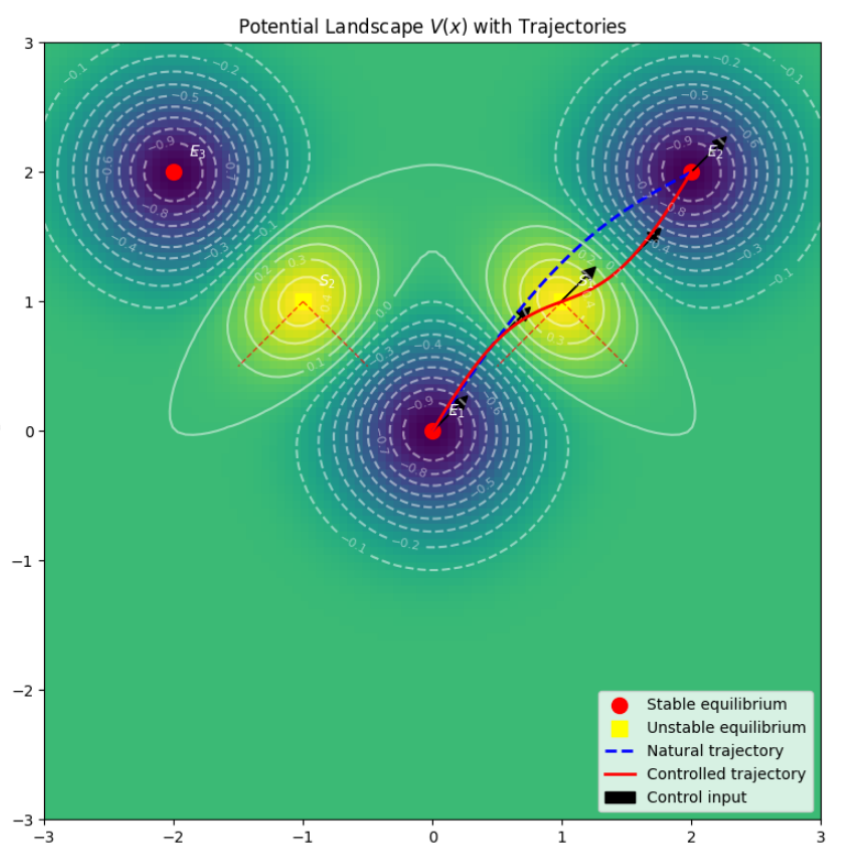 |
📊 Optimal Multi-Drug Dosage Control For Cellular Transitions *(Private Repo)*
Optimal Control Theory for Cell State Space Transition and Trajectory Planning Optimal control models to optimize biological state transitions and drug dosage planning. Manuscript-stage; not yet publicly released. |
 |
📊 TRANSFORMERS from a Mathematician's Lens. A mathematical perspective on attention and re-formalization of query-key-value parameterization. Manuscript-stage; not yet publicly released. |
⚙️ Advanced Transcriptomics & Spatial Omics
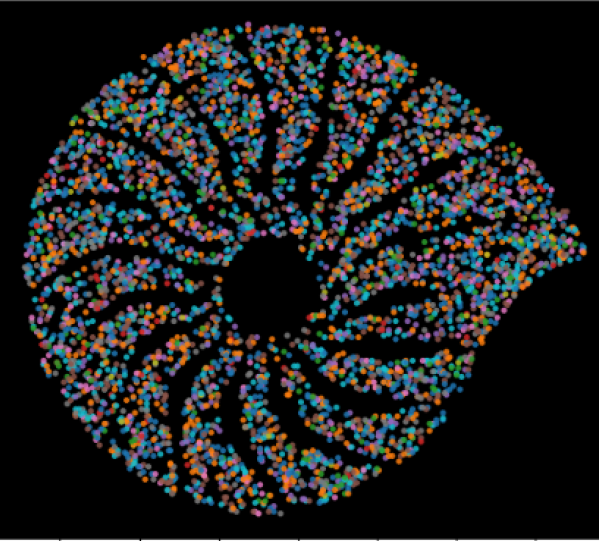 |
🔥 SPATIOME
Synthetic Platform for Advanced Transcriptomics Integrating Omics and Multidimensional Exploration. Computational framework for integrating high-dimensional transcriptomics data. GitHub Repository |
🚀 Kaggle AI Competitions in Biology With Interesting DATA:

|
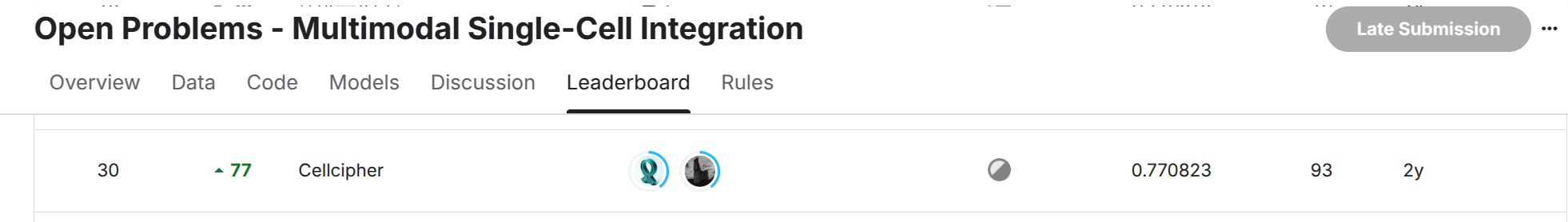 Private LB: Silver
Private LB: Silver
|

|
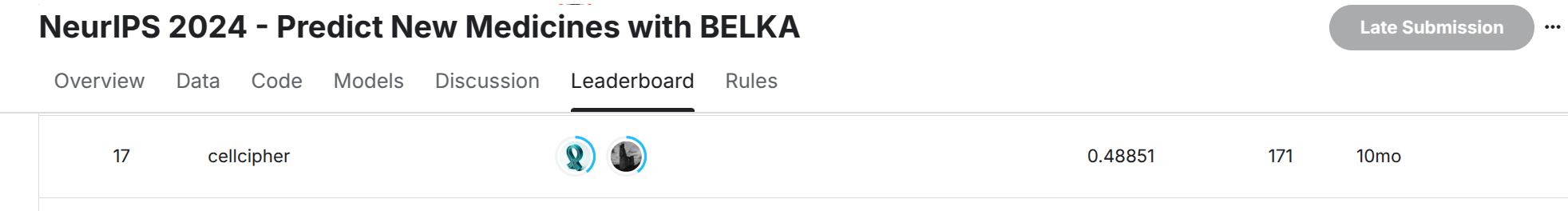 Public LB: Silver Public LB: Silver
|

|
| Kaggle is a Google Subsidiary |